> Range of assays > Genomic stability: Karyotyping assays > Duo iCS-Karyo
Duo iCS-Karyo:
combined G-Band and dPCR solution
Duo iCS-Karyo: the ultimate combined solution for hPSC genomic stability testing
Human Pluripotent Stem Cells (hPSCs) are particularly prone to developing abnormalities during their time in culture. Variants that appear can rapidly become predominant, compromising your research work.
Traditional G-Banding karyotyping is helpful in providing an exhaustive structural and numerical variant analysis. However, it does not have the required resolution to identify the most frequent genomic defects in human pluripotent stem cells.
G-Banding can detect defects down to 5-10 Mb. However, some recurrent hPSC abnormalities are smaller than that. This is the case for the gain of 20q.11.21 copy-number variant (CNV). The 20q.11.21 chromosomal amplification, commonly known as 20q, is detected in more than 20% of worldwide cultured human Pluripotent Stem Cells (hPSCs) (Avery et al., 2013; Halliwell et al., 2020) and represents 22.9% of the recurrent structural variants identified in these cell lines (Assou et al., 2020), making it the most common genomic abnormality in hPSCs.
This is why we recommend the combination of both G-banding and a more sensitive technique for a truly reliable genomic stability assessment.
2-in-1 solution: Get the exhaustivity of G-Banding analysis with the high-resolution of digital PCR
Duo iCS-Karyo combines both technologies to offer you the usual structural rearrangement analysis that the G-Banding karyotype offers, including balanced and unbalanced translocations, deletions, insertions and inversions.
Combined with iCS-digital™ PSC, our digital-based assay, it will also ensure that you catch over 93% of the most recurrent abnormalities found in hPSCs, such as the sub-karyotypic 20q.11.21 amplification.
Discover
Duo iCS-Karyo
in video
Watch a short presentation video explaining all there is to know about the latest genomic integrity solution by Stem Genomics called Duo iCS-Karyo. A 2-in-1 solution, combining iCS-digital™ PSC with G-Banding karyotyping, dedicated to the needs of research scientists working on hiPSCs.
Investigate Duo iCS-Karyo further: the importance of combining chromosome karyotyping and digital PCR assays to assess genomic integrity
This poster demonstrates the importance of combining methods to more robustly assess the genomic stability of pluripotent stem cells. Click on the image to understand more about the powerful combination of G-banding analysis and Stem Genomics iCS-Digital™ PSC to provide a comprehensive genomic stability assessment. This document demonstrates how one assay complements the other and delivers more robustness.
A comprehensive service to make your life easier
We provide full processing services, from metaphase chromosome preparation and DNA extraction to the final report.
The final analysis will be sent to you by email within 10 to 20 working days* of receipt of your live cells. It will consist of fully interpreted reports with representative images and a summary of the analysis.
You will also be able to count on our team of PSC genomic analysis experts for any queries you may have throughout the process.
Duo iCS-Karyo key specifications at a glance
Live cells
Acquisition of a new cell line
Banking characterization
End of workflow
Europe:
2 x T25 flasks at 60% confluency
+
50 mL of media
USA:
1 x T25 flask at 25% confluency
+
1 x T25 flask at 50% confluency
+ 50 mL of media.
Room temperature
iCS-digital™ PSC test:
– The most frequent abnormalities observed in hPSCs (CNVs) >20% mosaicism
– Sub-karyotypic abnormalities (>200bp)
Karyotype:
– Balanced and unbalanced translocations
– Aneuploidy
– Inversions
– Duplications / deletions
– Detect abnormalities >5-10 Mb and 10% mosaicism
10 to 20 working days from sample receipt*
* We always strive for fast deliveries but please note that lead times depend on planning and sample quality. Assuming the samples sent comply with the sample collection and shipment instructions, we can commit to a 10 business day lead time for the North-American market and a 20 business day lead time for Europe. Please note that we will not be able to plan the work on your cells without advance notice. Contact us at least one week before sending your samples to avoid disappointment.
For research use only. Available only in the European Union and North America. Outside these specific geographical areas, you can combine your G-Banding analysis with our iCS-digital™ PSC. Contact us directly for hESCs.
An accredited cytogenetician will conduct a 20-metaphase spread analysis for you in compliance with the European Guidelines for Constitutional Cytogenetics Analysis 2018 and the updated version of ISCN (International System for human Cytogenetic Nomenclature). For US karyotypes, work will be performed in the spirit of GLP (Good Laboratory Practice).
Available as a service only
(from one sample):
For research use only.
Practical testing guidelines
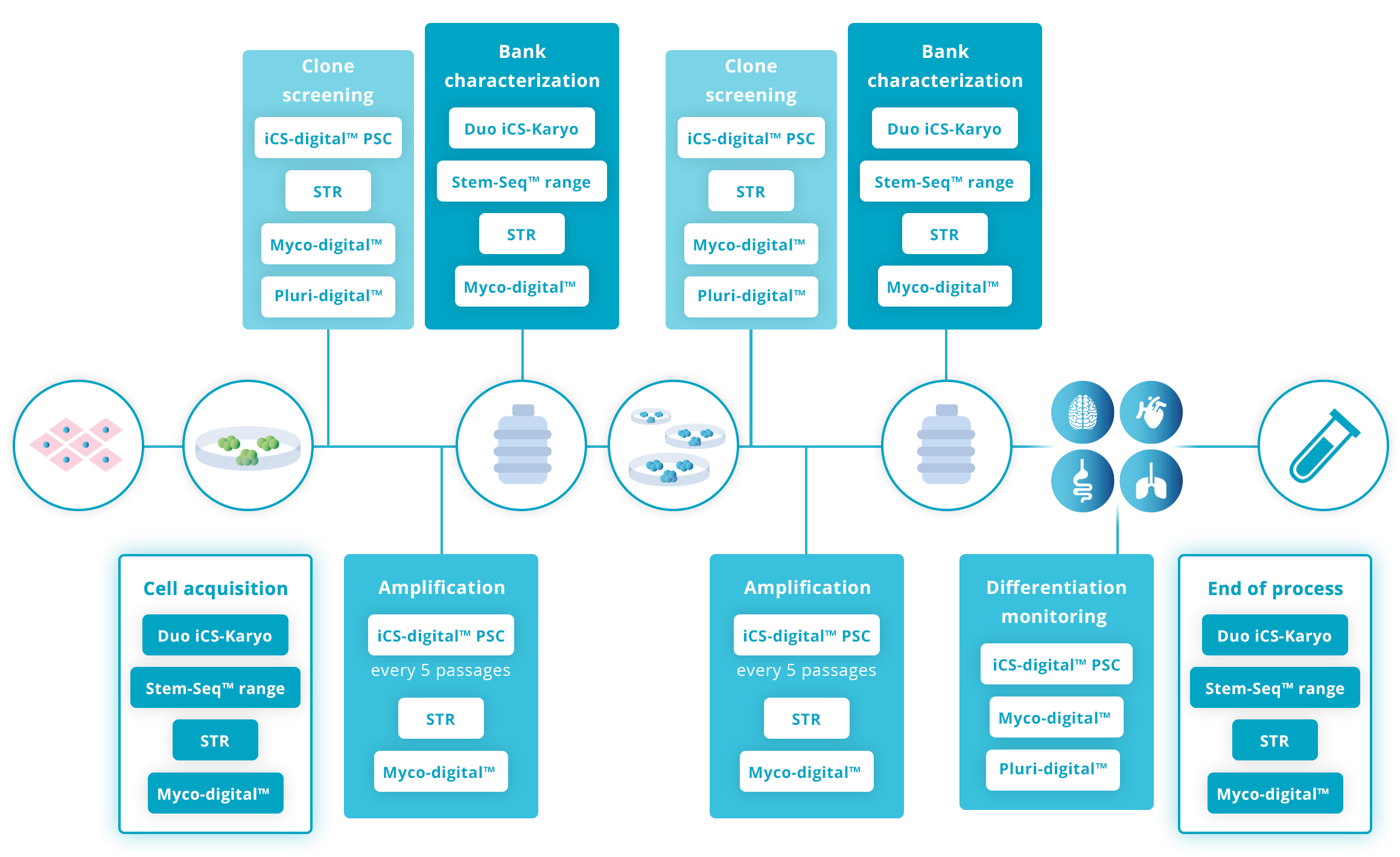
Recommended guidelines on genomic stability advise checking the cells every 5-10 passages during their time in culture as well as screening clones after any stressing process (reprogramming, gene editing, etc.) and during banking. It is also advisable when a new line is acquired.
Click on the graph for more details: